Action 6 – Monitoring of genetic diversity in Italian autochthonous breeds and its assessment
The action aims to monitor genetic diversity along generations in populations under conservation. Average inbreeding value within populations, calculated for the genotypes obtained by Action 2, will be compared to average inbreeding values obtained in the first TuBAvI project and the increase of inbreeding along generations will be studied (UniTO).
To see the results from the first TUBAvI project, click here.
Action 6 – TuBAvI (2017-2020): results
Monitoring of genetic diversity and inbreeding in Bionda Piemontese breed from 2016 to 2019 (UniTO)
The trend of genetic diversity and inbreeding, estimated by microsatellite molecular markers (Action 2), was monitored along generations in nucleus populations of Bionda Piemontese breed.
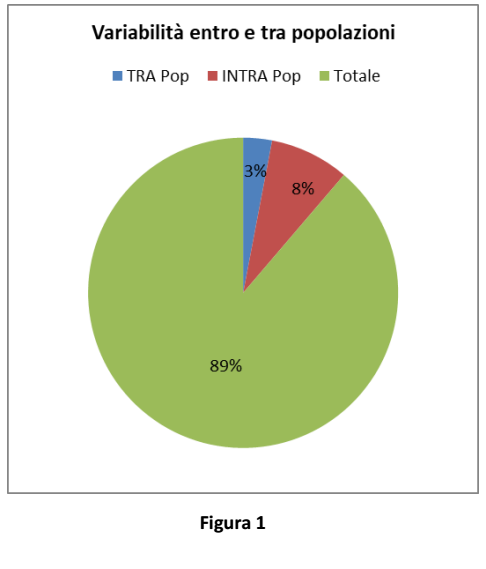
Figure 1 shows the result of the analysis of molecular variance along generations: low variance values among populations (made up of birds from different generations) mean that average heterozygosis remained comparable over time.
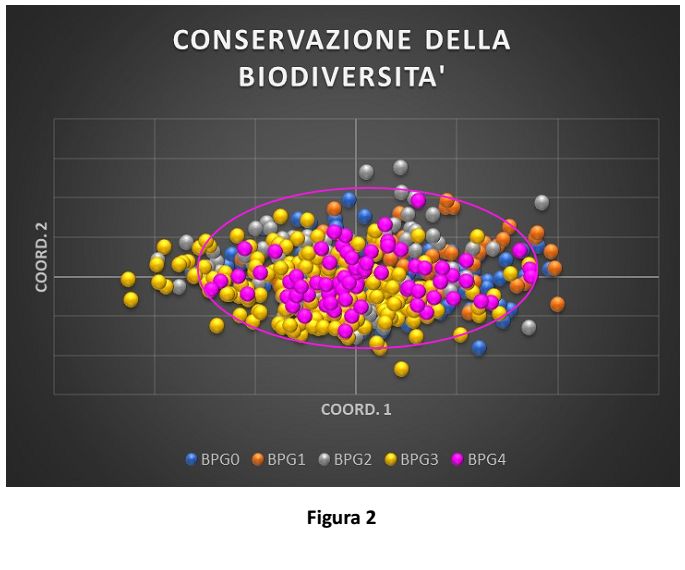
Figure 2 shows the distribution of birds on the basis of their genetic diversity (the more similar the birds are from the genetic point of view, the closer they are on the chart; the more different they are, the more distant they are in the chart). The surface covered by birds from the same generation shows the overall variability of the population (larger is the surface, and higher is the variability of the population). Each generation (from MPG0 to BPG4) is shown with a different color. The overlapping of different generations means that most variability is preserved. The ellipse surrounds the spatial distribution of the latter generation (BPG4).
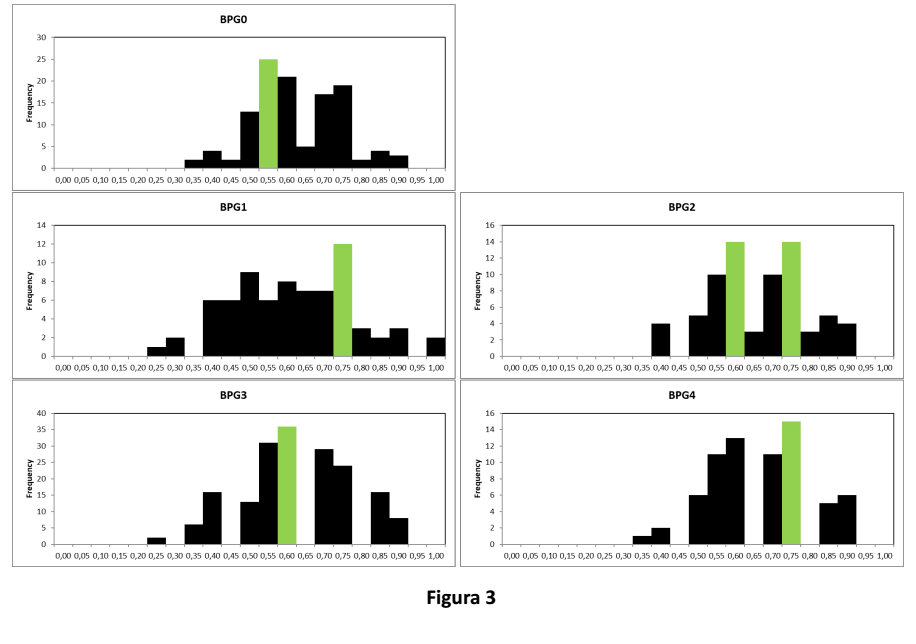
Figure 3 shows individual heterozygosis values (H-ind) recorded in birds from the same generation on the horizontal axis (from BPG0 to BPG4, similarly to Figure 2), and their frequences on the vertical axis. Values with highest frequence are shown in green. H-ind ranges from 0 (all loci are homozygous, no variability) to 1 (all loci are heterozygous, highest variability). Charts show that frequence of birds with lower H-ind values decreases over time, while frequence of birds with higher H-ind increases, thus showing an increase in individual genetic variability along generations.
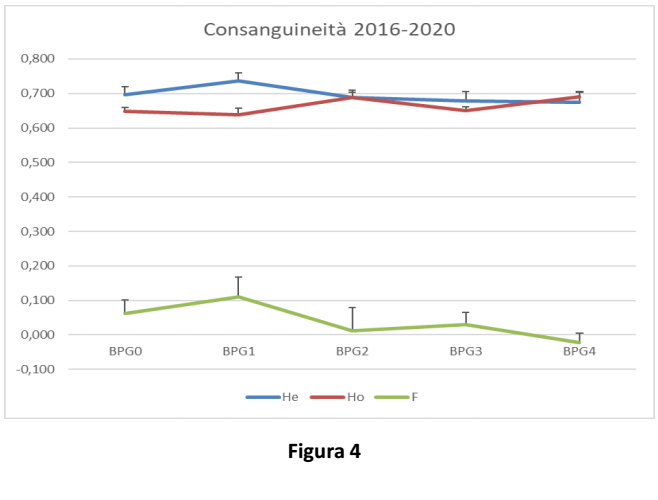
Nel grafico in Figura 4 è riportato l’andamento dell’eterozigosi attesa (He), dell’eterozigosi osservata (Ho) e della Consanguineità (F=indice di fissazione) nel corso delle generazioni: si può notare un rallentamento nella riduzione dell’eterozigosi attesa, contestualmente ad un aumento di eterozigosi osservata e di conseguenza ad una riduzione di consanguineità.
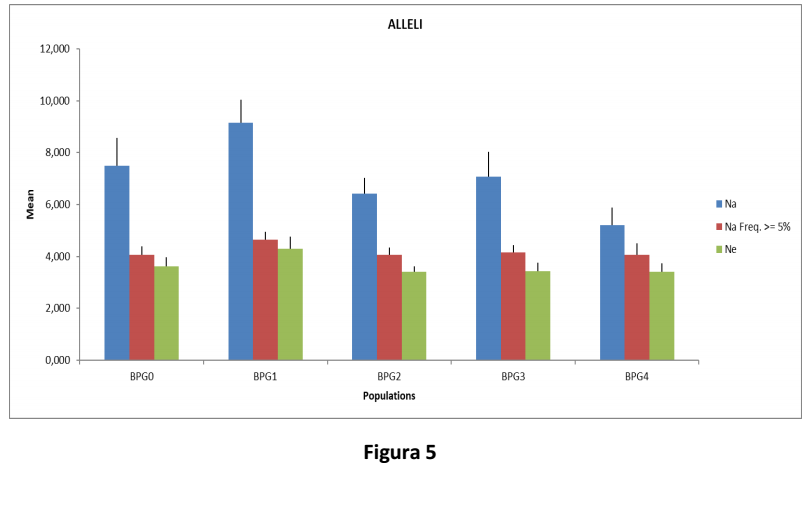
In Figura 5, il grafico riporta il numero di alleli osservato (Na) in ciascuna delle generazioni considerate. Il numero di alleli con una frequenza superiore al 5% (Na Freq. >=5%), così come il numero effettivo di alleli (Ne), si è mantenuto costante nelle ultime generazioni e quindi la variabilità allelica della popolazione è stata preservata.
Figure 4 shows the trend of expected heterozygosis (He), observed heterozygosis (Ho), and inbreeding (F=fixation index) along generations: reduction in expected heterozygosis slows down, while observed heterozygosis increases and, as a consequence, inbreeding decreases.
Figure 5 shows the number of observed alleles (Na) in each generation. The number of alleles with a frequence higher than 5% (NA freq >=5%), as well as the actual number of alleles (Ne), is constant along the latter generations, so allelic variability of the population is preserved.